HTIRC E-NEWSLETTER – May 2012
Volume 5 Issue 1
Articles in this issue:
- Website Provides Current Information to Thousand Cankers Disease
- Breakthroughs in New-Age Sequencing Technologies Could Accelerate HTIRC Research
- Butternut Restoration: Characterizing Disease Resistance and Site Preference
- Can Poplar Genes Function in Green Ash?
- Tropical Hardwood Tree Improvement and Regeneration Center Participates in Society of American Foresters National Convention
- Indiana Forestry and HTIRC Featured in Recent Public Television Special
- Recent Articles Highlight Greatwoods® Trees Planting and Care of Fine Hardwood Seedlings Publication Updates
Welcome to this issue of the Hardwood Tree Improvement and Regeneration Center E-newsletter. The HTIRC is committed to enhancing the productivity and quality of Central Hardwood Region trees and forests for the economic and environmental benefits they provide. Scientists at the HTIRC are using conventional tree improvement breeding as well as molecular and genetic technologies to improve the wood quality, growth characteristics, and insect and disease resistance of trees like black walnut, black cherry, red and white oaks, butternut and American chestnut.
Research in tissue culture, tree nursery practices, tree plantation establishment and management, and Central Hardwoods silvicultural systems is aimed at increasing the regeneration success rate for high quality hardwood trees and forests. Some interesting and unusual research areas include examining the potential for propagating trees with “figured” wood: birds-eye maple or curly walnut; and breeding trees that will be an economical source of bio-fuels.
Twice per year, we will attempt to provide interesting and useful information on Central Hardwood trees and forests, as well as sources for additional information and assistance.
Please pass this newsletter along to others who may enjoy or benefit from the information provided. If you would like a closer look at the HTIRC, please visit our web site at: http://www.htirc.org
Website Provides Current Information on Thousand Cankers Disease
The National Thousand Cankers Disease website, www.thousandcankers.com, was developed to help landowners, forest industry, resource managers, researchers, and other interested parties easily access information on Thousand Cankers Disease, which is currently impacting black walnut in Pennsylvania, Tennessee, Virginia, and several states in the Western United States.
Thousand Cankers Disease (TCD) was first identified in the Western United States on planted black walnut dying in urban areas, although subsequent evidence indicated TCD has been killing black walnut in western states for several years prior to positive identification. In 2010, dead and dying black walnut trees with TCD were identified in and near Knoxville, Tennessee. Subsequently, TCD was identified in the Richmond, Virginia area and Bucks County, Pennsylvania. These eastern US infections are suspected to be caused by walnut logs or walnut lumber with bark attached transported from infected western states. Quarantines have been established by several states to prevent the introduction of infested materials from known TCD infested areas.
If you would like to learn more about how to recognize TCD, what is currently being done to understand and combat this disease, and details on quarantines, please visit the National Thousand Cankers Disease website at www.thousandcankers.com .
This website is a collaborative effort between the Northeastern Area State and Private Forestry, the USDA Forest Service Northern Research Station, the Purdue University Department of Forestry and Natural Resources, the Hardwood Tree Improvement and Regeneration Center, and the Walnut Council.
On April 26 a meeting entitled Thousand Cankers Disease: Methods and Research and Technology Development Needs Assessment Workshop was held at Purdue University, West Lafayette, Indiana to bring together researchers, resource managers, landowners, forest industry members, and others interested in the black walnut resource to hear the most recent developments related to the disease and help develop a strategic direction for future research and technology development to address this threat.
This meeting was supported by the USDA Forest Service State & Private Forestry, the Hardwood Tree Improvement & Regeneration Center at Purdue University, the Walnut Council, and the American Walnut Manufacturers Association
Back to the Top
Breakthroughs in New-Age Sequencing Technologies Could Accelerate HTIRC Research
By Shaneka Lawson, Plant Physiologist
The use of advanced genetic and genomic techniques in tree research has helped researchers worldwide sidestep a number of hurdles that have slowed the progress of fine hardwood research. The continued need for the support of hardwood conservation has encouraged a number of groups to develop research programs designed to provide a greater array of resource data for hardwood research scientists.
In Europe, several groups are working to incorporate newer methods of analyzing oak (Quercus spp.) data for easier online access. In the United States, work on tulip poplar (Liriodendron tulipifera), American beech (Fagus grandifolia), American chestnut (Castanea. dentata), American sweetgum (Liquidambar styraciflua), Chinese chestnut (C. mollissima), northern red oak (Q. rubra) and white oak (Q. alba) has been the subject of several research projects aimed at using Next Generation Sequencing (NGS) technology for population mapping, expressed sequence tag (EST) sequence analysis; where a short piece of cDNA is used to annotate and identify genes, cDNA library construction and comparative sequence analysis. NGS allows millions of base pairs to be sequencing at a time compared to other, more traditional methods. As technological breakthroughs continue to happen genetic research companies each struggle to keep up with the promises they have made to investors and the media about the abilities of their particular technologies.
During a recent bioinformatics seminar at Purdue, it was confirmed that the recent acquisition of new NGS platforms by the Purdue University Bioinformatics Core Facility will offer a cost-effective method for conducting research (Figure 1). More time will be spent during data analysis phase of the process than ever before however the information obtained will be greater than ten times more informative (Figure 2). Older technologies that were previously available did not offer the sensitivity of NGS. As more time is spent in front of the computer lab spaces can also become more efficient and streamlined in response to the shift in research focus.
Within the Hardwood Tree Improvement and Regeneration Center (HTIRC) and its Tropical counterpart (TropHTIRC), a select group of scientists are crossing over into the realm of NGS to solve a myriad of problems in fine hardwoods research. This “crossing over” will allow the HTIRC and TropHTIRC research centers to stay on the very cusp of newly discovered and implemented technologies. NGS will also allow the scientists connected to the varied key focus projects of the centers to concentrate their efforts on fine hardwood species targeted for improvement rather than to wait until a particular genome is published by another group. Here at the HTIRC and TropHTIRC, the species of interest are koa (Acacia koa), black cherry (Prunus serotina), black walnut (Juglans nigra), English walnut (J. regia) butternut (J. cinera), American chestnut (Castanea dentata), ash (Fraxinus spp.) and oak.
A few of the proposed projects using NGS are centered around koa research while another is focused on black walnut. The principal investigators for these projects are Lawson, Michler, and Woeste. The projects are still in the early stages at this point and little information is available for dissemination however we are on the very edge of our seats waiting to see what is going to happen next. The HTIRC and TropHTIRC centers have moved forward by leaps and bounds during the last few years and seemed poised for bigger and better things in the coming years.
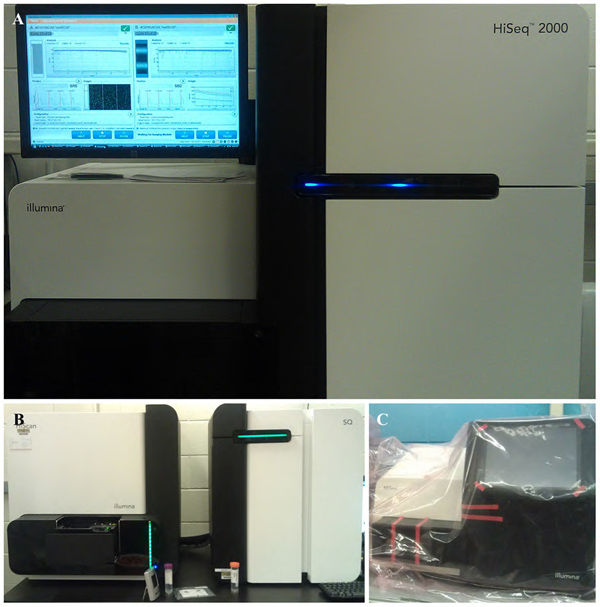

Figure 1. Views of the Next Generation Sequencing (NGS) platforms at Purdue University. The (A) HiSeq 2000 (B) HiScan and (C) MiSeq are three cutting edge machines capable of processing billions of bases within 2 weeks.
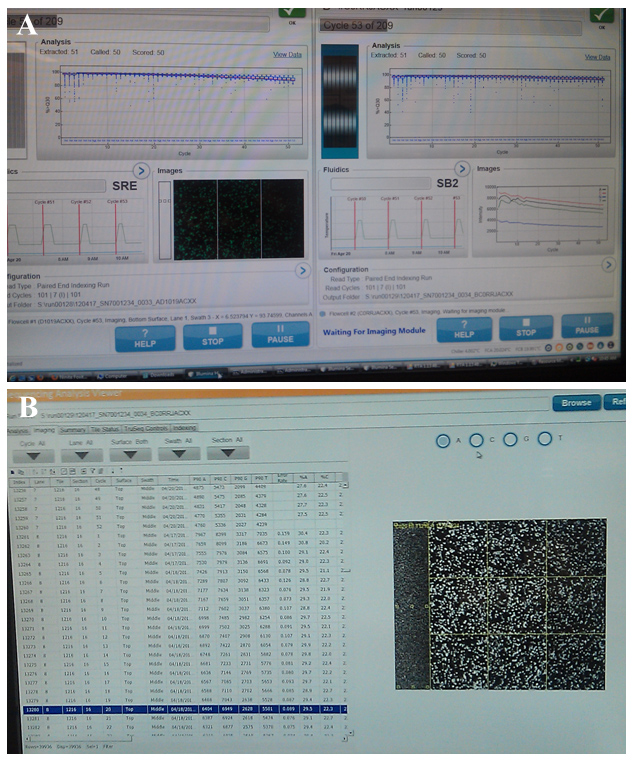
Figure 2. Images of an ongoing sequencing run using Illumina technology. After reading billions of nucleotide bases, the (A) data analysis and (B) annotation process which follows will often take months to complete.
Butternut Restoration: Characterizing Disease Resistance and Site Preference
By Phillip A. Crystal, M.S. HTIRC Graduate Student and James J. Jacobs, Ph.D. HTIRC Graduate Student
Butternut (Juglans cinerea L.) was once prized ecologically for highly nutritious nuts and economically in the forest products trade as “white walnut.” Over the past 50 years, populations of butternut throughout its native range have been decimated by the introduced pathogen Ophiognomonia clavigignenti-juglandacearum (OCJ; formerly Sirococcus clavigignenti-juglandacearum), cause of butternut canker disease. Observations of disease progress in forest stands have noted apparently healthy butternuts surrounded by diseased individuals. These trees were presumed to be either escapees (uninfected simply by chance) or trees that expressed some form of disease resistance. Scion material (branch segments for clonal propagation) and/or seed were collected to preserve these potentially-resistant trees for further study and are currently planted at genetic repositories throughout the United States. Solving the disease issue, however, will not restore butternut to the landscape. In addition to disease-related mortality, the species’ short life-span (70 years), lack of disturbance in eastern forests, and hybridization with Japanese walnut (J. ailantifolia), a nut cultivar introduced in the late 1800s, are all interacting to further reduce butternut populations. As a result, artificial regeneration will likely be required to restore the species across the landscape. Currently the HTIRC is simultaneously addressing two major impediments to restoration success: identifying disease resistant material and proper site conditions for outplanting butternut.
Butternut Canker Resistance Research
The ongoing search for resistance to butternut canker is currently limited by inadequate techniques to experimentally separate resistant trees from those susceptible and lack of understanding regarding the basic biology of tissue colonization by the pathogen. Ideally, potentially-resistant material would be rapidly-screened for their ability to resist disease development, allowing resources to be properly allocated toward breeding programs, effective seed orchard establishment, and eventual deployment of resistant trees onto the landscape.
Butternut canker is primarily a disease affecting areas of the tree where the stem has been wounded in some way, and current inoculation methods are tailored to mimic the most likely course of infection observed in nature. Wound inoculations use a device to penetrate through the outer bark and phloem, giving access to the tree’s cambium. The fungus, which has been grown on special growth-media in the lab, is then placed inside the wound and the area is wrapped to keep the wound from drying out. After a specified amount of time the wrapping is removed and disease progress is assessed. So far, the method of using wound inoculations to separate resistant and susceptible trees has produced inconclusive results; for example, many trees that were apparently canker-free under natural conditions are readily colonized by the fungus after artificial inoculation. Further complicating the disease screening methodology are the multiple generations of hybrid progeny between butternut and closely-related, Japanese walnut. Many of these hybrid trees appear to be functionally resistant and some of the original “resistant butternuts” have turned out to be hybrids.
There are many possible explanations for the differences we have observed between the resistance of trees under natural conditions (in the field) and their level of resistance in response to artificial infection. The two most likely explanations are related to our inoculation methods. (1) The amount of fungus we used in the artificial inoculations may have been too high (unnaturally high), allowing the fungus to overwhelm the tree’s defenses and permitting the fungus to overcome potential resistance mechanisms present in the selections. (2) The observed natural resistance results from constitutive resistance mechanisms (those not induced by pathogen colonization) present in the outer bark and phloem. Scientists at HTIRC are refining their method of field inoculation to more closely mimic what is observed in the field so that inoculations are uniformly successful on trees known to be susceptible. This will enable researchers to better distinguish resistant from susceptible butternut families.
Recently, OCJ was found actively infecting leaves of a butternut and a hybrid. It is possible symptom development on leaves will reflect total tree tolerance/resistance to the pathogen. To test this possibility, a detached leaf disease screening assay was developed during the 2011 growing season.(Figure 1) The development of stem canker inoculations made during fall 2011 will be correlated to foliar lesion development during spring 2012. If the infection in a detached leaf assay is well-correlated with stem canker development of individual trees, butternut disease screening efficiency will be markedly increased compared to the wound inoculation method described above.

Fig. 1. Leaf assay of susceptibility of butternut, butternut hybrids, and black walnut to OCJ.
Photos take 7 days after spores of OCJ in water were placed on detached leaflets.
The development of butternut canker disease has never been thoroughly documented microscopically. Scientists at HTIRC have begun research to ascertain what, if any, disease resistance responses are present in the bark or phloem of healthy butternut trees. The goal is to observe if there are resistance responses that are bypassed when trees are artificially inoculated using the wounding method described earlier. Many tree species express disease resistance responses in their bark. Some of these response types, known as constitutive defense mechanisms, could explain the discrepancy between artificial inoculation success and rates of natural infection. If so, they should be considered in butternut breeding decisions.
Three separate technologies will be utilized to fully explore these constitutive defense mechanisms. Scanning electron microscopy will be used to characterize the interaction between OCJ spores and the outer surface of butternut tissues. We will also document fungal spore germination patterns on different butternut tissues. Light microscopy in conjunction with common histological stains will be used to characterize resistant and susceptible reactions following inoculation. Next, to more specifically track the development of OCJ in butternut tissues, we will attempt to genetically transform the canker fungus with a Green Fluorescence Protein (GFP) gene. This gene, originally isolated from a marine jellyfish, produces a protein that, under controlled laboratory conditions, will cause the fine strands of fungus growing in butternut tissues to glow green when exposed to ultraviolet light. This technology will allow quick visual separation of the fungus from surrounding tree tissue, so we can monitor in detail the course of an infection and the tree’s response.
Surprisingly, healthy trees are like living hotels. Large numbers of microorganisms live on and in them without causing any obvious symptoms. In fact, some of these microbes, which are called endophytes (live inside) and epiphytes (live outside), may influence the presence or absence of disease or significantly inhibit the growth of OCJ. The relationships between these microorganisms and the development of disease by an outside pathogen are interesting for many reasons, including their potential as biocontrol agents. Several forms of “biocontrol” are possible as a result of this research project: breeding butternut to preferentially accept colonization by certain endophytes, direct application of an endophytic or epiphytic organism in areas of butternut’s range where it may not be found at sufficient population levels, or if one of these organisms is able to directly inhibit growth of OCJ, the chemical basis for this inhibition can be isolated and directly applied as a tree protectant. We have collected fungi from butternut, Japanese walnut, and their hybrid growing on two different sites.(Figure 2) Hundreds of fungal endophytes and epiphytes have been isolated and we are in the process of identifying those using morphological and molecular techniques. Following their identification, representative samples will be tested for their ability to inhibit the growth of OCJ in culture.(Figure 3)

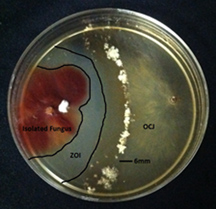
Figure 2. Sample of fungi isolated from butternut, hybrid and Japanese walnut tissues during Fall 2011 collection.
Figure 3. A fungal endophyte (red-colored) of butternut can inhibit and prevent the growth of OCJ in vitro. The Zone of Inhibition (ZOI, clear in the image on the right) forms because growth of OCJ is affected by the presence of the endophyte.
Basic Research in Butternut Afforestation
Lack of information about site preference and lack of management guidelines hampers the successful deployment of butternut seeds and seedlings. The literature on butternut silviculture is particularly sparse, but it may expand rapidly if we can determine that butternut has requirements similar to its well-studied relative, black walnut (J. nigra L.). Both butternut and black walnut have high light and nutrient requirements, however, information on butternut’s moisture tolerance is conflicting. Observational accounts indicate butternut prefers streambank sites, but also tolerates dry, high pH soils better than black walnut, despite butternut’s shallower root system. While some tree species are site generalists, studies including butternut report poor performance across a wide range of conditions.
The site preference issue is further complicated by the presence of the morphologically-similar hybrids between butternut and Japanese walnut. These are unknowingly included in butternut plantings or labeled as “butternut hybrids,” implying that the hybrids are similar to butternut in their growth habit and ecological attributes. While no information exists on site preference of hybrids, Japanese walnut is distributed throughout riparian and floodplain forests in Japan, indicating butternut and hybrids may both prefer wet sites.
To assess moisture tolerance and evaluate hybrids, black walnut, butternut, hybrids, and Japanese walnut seedlings were exposed to drought, flood, and control conditions in the greenhouse during the 2011 growing season. Flooded butternut, droughted Japanese walnut, and hybrids under both stress conditions had reduced photosynthetic assimilation (a non-destructive measure of stress) and leaf area. Results indicated that the butternut families tested were flood intolerant, implying that butternuts should not be planted in poorly-drained areas. This finding was not inconsistent with the observation that butternut grows along stream banks. Bank positions are extremely variable in moisture stress, with low sites prone to flood and high sites prone to drought. Our results indicated that hybrids were tolerant of a much narrower range of site conditions than either parental species. This could have been the result of using naturally-occurring hybrid families, some of which genetic testing revealed to have greater proportions of Japanese walnut to butternut DNA, and vice-versa. Hybrids to be used in the conservation effort should be carefully selected for butternut character. Yet, this suggestion is currently only an ideal as there is no consistent way to distinguish hybrid and pure butternut seedlings on appearance alone, and genetic testing is both expensive and time consuming.
Because all the seedlings we used in the stress tolerance studies were root-wrenched in the nursery, these results are not fully generalizable to natural regeneration (trees grown from seed). Root wrenching alters the root (water uptake) to shoot (water loss) ratio by ripping deeper roots off and promoting fine root formation, which allows for easier replanting and increases establishment success. To better assess the ecological differences between butternut and hybrids and to identify physical traits that consistently differ between them, butternut, Japanese walnut, hybrids, and black walnut families will be grown from seed during the 2012 growing season. This will preserve their root architecture and root to shoot ratio for analysis of moisture preference and allow us to evaluate discrepancies in competitiveness. In addition, the HTIRC in conjunction with the College of Agriculture at Purdue University established in 2010 and 2011 seven test plantations of butternut, hybrids, northern red oak (Quercus rubra), and black walnut across a range of site conditions including: slope position (bottomland to upland), soil texture, and depth to water-table. As these plantations mature they will provide much needed information on butternut site preference and the potential role of hybrids in butternut restoration.
For more information about butternut, visit the HTIRC’s homepage (www.htirc.org) and click “Butternut Links.” We highly recommend Dr. Keith Woeste’s 2011 Butternut Newsletter on that page, as it describes a potentially delicious way (think butternut cookies) you can help restore the species and additional information on the state of restoration efforts. Other great sources of knowledge include the U.S. Forest Service Silvics Manual and Purdue Extension Publications: FNR-420 and 421.
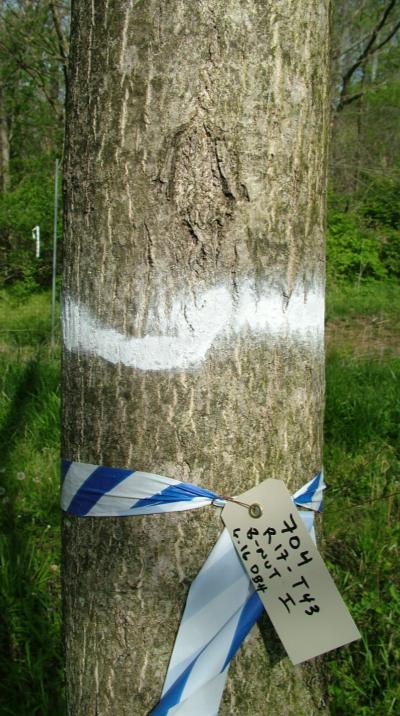


The butternut on the left is showing common symptoms of early butternut canker infection, with dark oozing cankers and gradual death of cambium. The butternut on the right was intentionally infected with butternut canker where the small scar exists on the upper middle portion of the trunk. This tree was able to resist expansion of the disease, indicating potential resistance to butternut canker-caused damage.
Can Poplar Genes Function in Green Ash?
Green ash (Fraxinus pennsylvancia Marsh.) is a hardy, fast-growing hardwood tree native to the mid-west and eastern United States as well as portions of Canada (Figure 1). This tree has been the popular choice of growers for decades because of its high value wood and robust growth. Green ash trees have also been used in urban renewal projects because of its superior growth form and inherent tolerance to adverse environmental conditions. Poplar, (Populus tremula x P. alba (717-1B4) in particular), another tree with robust growth, is one of the most widely used gene study model tree species in the world. Since the sequencing of the Populus trichocarpa genome in 2006, researchers have continued to use gene information from this tree to identify how various morphological or physiological traits could be altered by the overexpression (a method of artificially increasing the expression level of a gene) of a single or multiple genes.
The insertion of poplar DNA into green ash required genetic transformation. Transformations of green ash were conducted using an optimized green ash transformation protocol published by Du and Pijut (2009). The use of this protocol resulted in a confirmed transformation rate of greater than 4% (Figure 2). A total of 14 green ash trees overexpressing a poplar stomatal density gene were generated and confirmed using polymerase chain reaction (PCR). All plantlets were initially grown in aseptic culture conditions before being acclimatized and transferred to the greenhouse.
In this study, a poplar gene recently shown to function in stomatal density and patterning was overexpressed in green ash trees. The resulting green ash transgenic plants showed variations in stomatal density and patterning. Compilations of stomatal density data taken from the 14 transgenic green ash trees indicated that stomatal densities were extremely varied in the generated plants with the greatest variation observed being 31% lower than untreated plants. However, closer inspection of the transgenic green ash leaves indicated that stomatal distributions across the leaf surface were altered as well. Examination of the same regions on the leaves of both the untreated and transgenic plants eliminated the potential influence of location on stomatal differences as shown in this example using poplar (Figure 3). The stomata of transgenic green ash plants were distributed in small groups rather than in a fairly uniform pattern (Figure 4). These data indicated that the poplar stomatal density gene served a similar function in stomatal density when overexpressed but also caused an unexpected stomatal distribution physical characteristic.
Continued study on these transgenic plants could yield many other characteristics to demonstrate that the overexpression of a poplar gene in green ash tree may not result in responses identical to those obtained in poplar. This finding is preliminary but does show the possible overlap in gene function between two different tree species. Integration of DNA (via transformation) into a species that has not been fully sequenced is a process that merits further study. It is likely that greater gains will be achieved once the green ash genome has been fully sequenced.
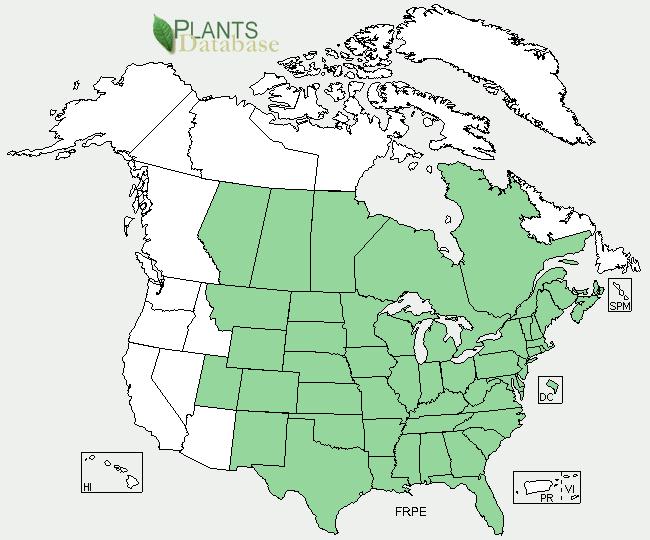
Figure 1. Green ash range. The extended growth ranges and adaptability to various climate conditions has been a key factor in the ability of green ash trees to establish stable breeding populations in most Canadian provinces and all of the lower 48 states east of Idaho, Nevada and Arizona. Image from the USDA Natural Resource Conservation Service Plants Database (http://plants.usda.gov/java/).

Figure 2. Green ash transformation. Several thousand green ash seeds were sterilized before the hypocotyls were extracted for transformation. Potential transgenic plants were placed on selection medium following transformation to determine survival rate before being transferred to a medium which promoted elongation into plantlets.
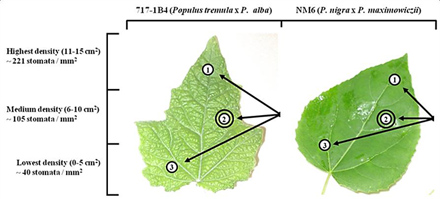

Figure 3. Stomatal sampling using poplar leaves. Visualization of 717-1B4 sampling regions to demonstrate that stomatal densities vary dependent upon sampling location. The region used in this study was region 2 (double circle). Densities given were from a sample leaf from a wild-type 717-1B4 plant.
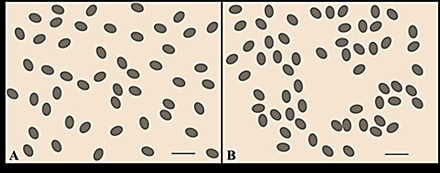
Figure 4. Green ash stomatal clustering. A comparison of control green ash trees and a sample transgenic line with similar stomatal density indicated that the stomata of (A) control trees were more evenly distributed while those of (B) transgenic trees displayed a more compact, clustered arrangement. Bar = 0.05 mm
References
Du, Ningxia; Pijut, Paula M. 2009. Agrobacterium-mediated transformation of Fraxinus pennsylvanica hypocotyls and plant regeneration. Plant Cell Reports. 28: 915-923.
Tropical Hardwood Tree Improvement and Regeneration Center Participates in Society of American Foresters National Convention
The Society of American Foresters, the largest professional association for foresters in the United States, had their national convention in Honolulu, Hawai’i in November, 2011 and the Tropical Hardwood Tree Improvement and Regeneration Center was there to welcome them. The Tropical HTIRC was represented by a booth in the exhibit hall where four Hawaiian hardwood bowls crafted by some of the best wood turners on the islands were raffled and raised over $1700 for the SAF Science Fund. Three other magnificent bowls were on display and now reside at the Hardwood Tree Improvement and Regeneration Center at Purdue University. Research projects associated with the Tropical HTIRC were part of the Silviculture Instructors Tour on the Big Island prior to the convention, and several partners associated with the Center provided research presentations as part of the scientific and technical concurrent sessions presented at the convention.
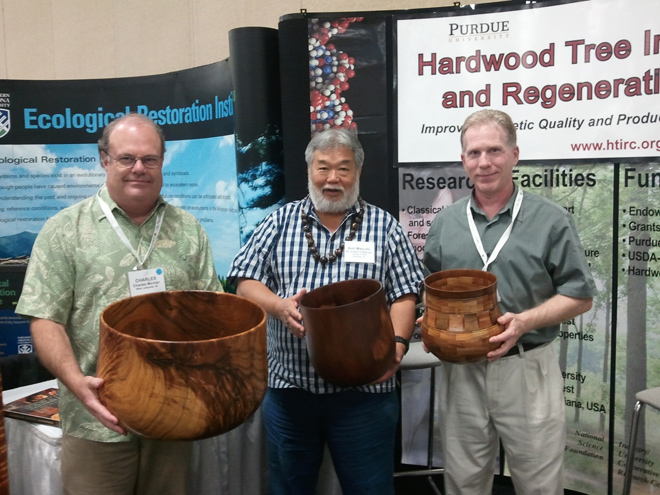

(Charles Michler, Bob Masuda, and Rob Swihart pictured)
Three Hawaiian hardwood turned bowls exhibited at the SAF national convention in Honolulu. The bowls are, from left to right, Koa held by Charles Michler, Tropical HTIRC director, Milo held by Bob Masuda, Outreach Coordinator for the Tropical HTIRC, and a segmented bowl made of Koa, Milo, Kou, and Kamani, held by Rob Swihart, Department Head for the Purdue University Department of Forestry and Natural Resources.
Indiana Forestry and HTIRC Featured in Recent Public Television Special
Hardwood Tree Improvement Regeneration Center staff and research projects were part of a recent public television special entitled Indiana Expeditions – Forests at Work. Indiana Expeditions is an award winning educational series originating from WFYI Indianapolis Public Television and is hosted by Rick Crosslin. Indiana Expeditions – Forests at Work presents a broad look at Indiana forests, forest products, and the people who help manage the forests of Indiana in a special one hour episode recently airing on Public Television.
You can see segments or watch the entire episode at: http://www.wfyi.org/IndianaExpeditions/ForestsAtWork.asp
Recent Articles Highlight Greatwoods® Trees
The Hardwood Tree Improvement and Regeneration Center is working to improve the growth, quality, and disease resistance of several fine hardwood tree species, including black walnut, butternut, oaks, black cherry, chestnut, and most recently Koa, a highly valuable native Hawaiian hardwood. Some of these improved trees are available for licensing under the Greatwoods® trademark, as announced through a variety of media outlets recently. If you would like to see the press release article on Greatwoods you can visit this website:
A web video highlighting the Greatwoods® trademarked trees and HTIRC research activities can be found at this website: http://youtu.be/9P6At_PfhL0
Planting and Care of Fine Hardwood Seedlings Publication Updates
Landowners and natural resource managers have a collection of publications entitled Planting and Care of Fine Hardwood Seedlings available through the Hardwood Tree Improvement and Regeneration Center, providing information on establishment and management of fine hardwood trees. One of those publications, Financial and Tax Aspects of Tree Planting, was recently updated to reflect changes in the tax code related to tree planting activities.
You can access all the publications in the series from the HTRIC website at www.htirc.org
Have questions about tree planting? This series of publications can be viewed or downloaded free of charge. Planting and Care of Fine Hardwood Seedlings
Van Eck Scholarships available for graduate research with the HTIRC.
Ask the HTIRC: email Lenny Farlee with your tree planting and forest management questions and we’ll help you find the answers.

